



Antimicrobial Resistance – Veterinary and Public Health Concerns in Europe
Radical reforms of antimicrobial use in livestock are being contemplated for political reasons, according to David Burch of Octagon Services Ltd.Antimicrobial resistance in animals and its potential to spread to man has become one of the hot issues involving scientists, regulators and politicians in Europe. What to do about it seems to be the main subject; should we ban certain drugs (Danish approach), ban in-feed antimicrobial use (Dutch approach), should we limit their availability and distribution or ban them completely in animals? Currently, this is under consideration by the European Commission (DG Sanco). In the meantime, each country is adopting or has adopted its own approach to managing the problem.
Dr Burch expresses his concern that radical reforms are being contemplated for political reasons, when even the latest report from EFSA BIOHAZ (2011) committee states that the establishment of risk factors is complicated by ‘data unavailability’ or the ‘lack of its accuracy’. This sounds like the same confusion over the banning of antimicrobial growth promoters on the basis of the ‘precautionary principle’, when really, by now, we should be making informed decisions on a good scientific basis.
Antimicrobial Resistance Development
Antimicrobial resistance and its development is a complex subject. Originally, it was a natural defence mechanism of bacteria to fight against naturally occurring antibiotics, which are produced by a number of fungi and bacteria in the wild, to aid their survival. Now, we are artificially exposing them to antibiotics as we treat animals, to fight disease.
Antibiotics are produced commercially by fermenting these fungi and bacteria and extracting the antibiotic (e.g. penicillins, tetracyclines, aminoglycosides) or frequently adding side chains and producing semi-synthetic antibiotics (e.g. amoxycillin, methicillin, cephalosporins) with different or improved spectra of activity, pharmacokinetics or efficacy. There are also fully synthesised antimicrobial compounds, like the sulphonamides, trimethoprim and the fluoroquinolones. Each antimicrobial family and its subgroups has its own mode of action and thereby each bacterial or mycoplasmal species develops its own way of countering the antimicrobial, as a defence or resistance mechanism, so that it can survive and continue to live in the environment it inhabits, e.g. the gut, the respiratory tract etc. The main antimicrobial families, their mode of action and common resistance mechanisms are summarised in Table 1.

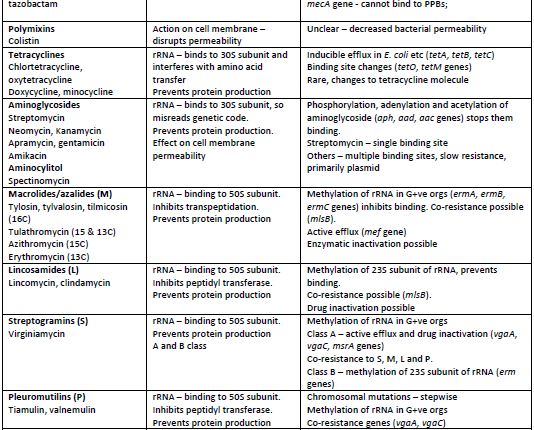
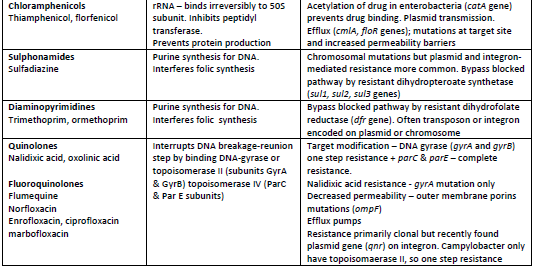
Simply, the bacterium is constructed of an outer cell wall of variable thickness with an inner cell membrane. It has chromosomal DNA in a tightly coiled chain, which controls growth and multiplication. The DNA sends messages to the ribosome (rRNA 50S subunit and 30S subunit) via messenger RNA (mRNA) to produce polypeptides or proteins for growth. Transfer RNA (tRNA) carries the amino acids to the ribosome to form the new proteins. When the bacterium is ready to divide the DNA uncoils and divides and a new bacterial cell is formed. Some bacteria multiply rapidly, like E. coli and some grow slowly like Brachyspira spp. The rapid, prolific growers have more of a chance to develop new DNA mutants and these mutations may increase resistance to antibiotics. All the bacterial structures can be targets for antimicrobial attack. The penicillins or beta-lactam antibiotics target the cell wall; the polymixins, the cell membrane; the fluoroquinolones, the DNA, and the tetracyclines, macrolides, pleuromutilins, aminoglycosides target the RNA.
Bacteria are routinely classified as Gram positive (blue staining with Gram stain – due to a thick cell wall) these include Staphylococcus spp, Streptococcus spp, Enterococcus spp and Clostridium spp. Gram-negative (pink–staining – thin cell wall) bacteria are primarily found in the gut, such as E. coli, Salmonella spp, or in the respiratory tract Actinobacillus pleuropneumoniae, Pasteurella multocida and Haemophilus parasuis.
They are further divided into aerobic (i.e. needing oxygen to survive) or anaerobic where they do not use oxygen and have different metabolic pathways. Enterococcus spp and Clostridium spp are examples of Gram +ve anaerobic bacteria and are found in the large intestine or colon and Brachyspira spp are examples of Gram –ve anaerobic bacteria, also found in the colon. Some bacteria can live in both environments, like E. coli. The commonly monitored bacteria for public health and regulatory resistance monitoring are the commensal bacteria, such as E. coli and Enterococcus spp, and Salmonella spp (mainly S. Typhimurium in pigs) and Campylobacter spp (mainly C. coli in pigs) for potential zoonotic infections, those infections in animals that cause disease in man.
Resistance Mechanisms
When we look at antimicrobial resistance, there are some other key factors to consider. Some bacteria are intrinsically resistant to certain antibiotics, usually due to their mode of action. For example penicillins, which act on the cell wall of a bacterium, are not effective against Mycoplasma spp, as they do not have a cell wall, only a cell membrane. Macrolides, like tylosin, cannot penetrate the cell membranes of certain Gram -ve bacteria like E. coli; aminoglycosides work poorly against anaerobic bacteria, as they use an oxygen-dependent mechanism to penetrate the bacteria. Susceptible bacteria can acquire resistance by a variety of mechanisms:
- Prevent an antimicrobial substance reaching a target by reducing its penetration into the bacterial cell often via porin changes, as they are often large molecules
- General or specific efflux pump mechanism to expel antimicrobial agents from the bacterial cell
- Antimicrobial agent inactivated by modification or degradation either before or after penetrating the cell
- Antimicrobial target may be modified so that it cannot act on it, or the microorganism’s activation or acquisition of an alternative pathway rendering the target dispensable (see Figure 1.)
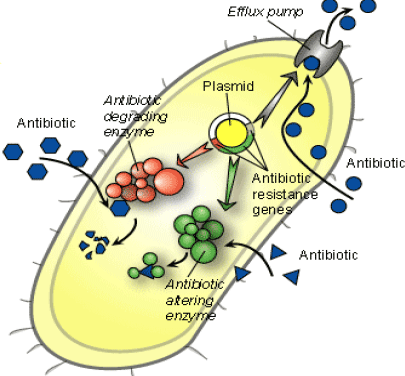
[Picture courtesy of A. Pridmore]
Acquired resistance can be achieved by a number of mechanisms, which are usually the result of selection pressure from the use of antibiotics. Mutations in the chromosomal DNA, which then alter the DNA coiling etc, are important for the fluoroquinolones. DNA changes which affects the binding sites of the ribosome are important for the macrolides, lincosamides, streptogramins and pleuromutilins and co-resistance can occur between these families, as their sites of action often are close or overlap.
The acquisition of resistance genes from outside the bacterial cell is also highly important. Some bacteria pick up extraneous DNA genes from other broken down cells by transformation and insert them into the chromosome. Others receive DNA into the chromosome via transduction from viral bacteriophages but the most common route is via plasmid transfer at conjugation of two cells. The plasmid can be independent of the chromosome in the cell and made up of a variety of DNA genes or open-reading frames (ORFs), which may be significant or not. Plasmids can carry multiple-resistance genes, which are often carried in transposons or integrons, which are sections of genetic material that can insert themselves via enzymes transposases and integrases, respectively, usually into plasmids but also into the chromosome of a bacterium. This is a very common route of resistance transmission between enteric bacteria, like E. coli; hence they are good indicators for monitoring resistance.
MRSA – First Major Controversy
The penicillins or beta-lactam antibiotics have been the recent cause of concern following the discovery of methicillin-resistant Staphylococcus aureus (MRSA) infections, mainly clonal complex CC398, spreading in piggeries to the pig farmers and their families and also slaughterhouse workers and veterinarians. These were picked up originally in the Netherlands when farmers attended hospital and were screened for human MRSA. Fortunately in Man, it does not appear to have spread into the general population. It has been shown that this strain has spread across Europe possibly associated with pig movement (EFSA, 2010) down breeding pyramids and in growing pigs but also by selection from the use of advanced beta-lactams, such as the third–generation cephalosporins, which are becoming commonly used in some countries at processing (castration etc).
When originally used in Man, penicillin use soon caused resistance development, especially in Gram +ve infections such as staphylococci and they produced resistant penicillinases (beta-lactamases), enzymes that destroyed the penicillin molecule. Newer penicillins, which were beta-lactamase resistant, such as methicillin, were introduced. To survive, the staphylococci mutated and changed the enzymes in the cell wall, where the penicillins bound (penicillin-binding proteins – PPB). This was associated with the new mecA gene in the chromosome and this meant that none of the penicillins or cephalosporins was effective. Generally, the strains in pigs are different from the MRSA clones found in humans, particularly those associated with hospital treatment but it was a cause of concern, especially as there are few other drugs that the doctors can use to treat them.
ESBLs – Next Major Controversy
Ampicillin and amoxycillin were two extended or broad spectrum penicillins that treated Gram +ve and Gram -ve bacteria. As usage has increased, resistance has developed especially in E. coli, due to beta–lactamase production (see Figure 3). It was found that these could be blocked by the use of beta-lactamase inhibitors, such as clavulanic acid, which irreversibly bound to these enzymes and allowed the antibiotic to carry on working.
This combination approach is commonly used in veterinary and human medicine. However, what has caused some confusion is the term extended-spectrum beta-lactamases (ESBLs), as these refer to beta-lactamase enzymes that attack third- and fourth-generation cephalosporins. These are the next major controversy, as the genes can be transmitted via plasmids, relatively easily, amongst enteric bacteria such as E. coli and potentially Salmonella spp. These enzymes are usually susceptible to beta-lactamase inhibitors but some are developing resistance. Some cephalosporinase resistance genes, AmpC, are not susceptible to these inhibitors. These bacteria can be treated in human medicine by carbapenems but of considerable concern, carbapenemase-resistant bacteria are now being reported in human cases in Asia and have arrived in human medicine in Europe. Fortunately, these antibiotics are not used in veterinary medicine. Of significant interest, ESBLs associated with cephalosporin use were looked for in poultry, pigs and cattle (EFSA – BIOHAZ, 2011) in EU Member states (see Figure 2).
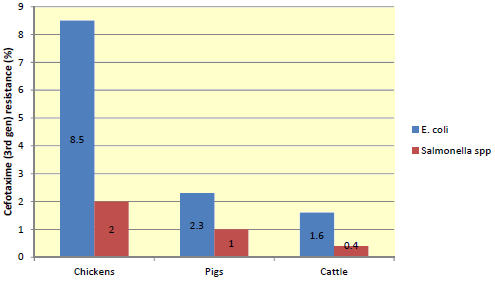
Surprisingly, the highest level of ESBL resistance was in chickens not pigs or cattle, but it was present at low levels. Apparently, chicks are injected in-ovo or at day–old and even if not used at broiler chick level, the resistant organisms have been shown to come down the breeding pyramid from imported birds into Sweden, as the drugs were not used there (SVARM 2010, 2011). Spain had the highest reported prevalence of ESBL resistance in E. coli from chickens at 26.4 per cent but only 1.1 per cent in pigs and 0.4 per cent in cattle. As the cephalosporins are not approved for use in chickens nor is there a MRL (maximum residue limit), perhaps a good method of control would be to stop their off-label use in poultry.
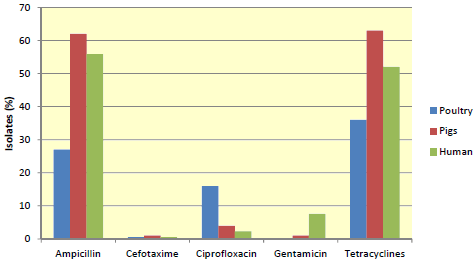
In a different EU survey (EFSA/ECDC), they compared antimicrobial resistance found in humans with that in animals for zoonotic bacteria Salmonella spp and Campylobacter spp. Salmonella enterica Typhimurium infections in Man are thought to be primarily associated with the consumption of pork (see Figure 3). Salmonella Enteritidis from poultry is still the major cause of salmonellosis in man.
In this case, resistance to ampicillin and the tetracyclines is high in pigs but to cefotaxime (3rd generation cephalosporin (ESBL resistance marker) it is very low. The fluoroquinolone, ciprofloxacin, resistance is also high in poultry but low in pigs. In this case, S. Typhimurium resistance patterns in man are generally similar to pig resistance patterns, suggesting that it is a significant contributor in comparison with poultry. Figure 3 also nicely demonstrates the lack of ESBL selection by other antibiotic use, other than cephalosporins.
In the case of Campylobacter coli, the resistance pattern is reversed (see Figure 4).
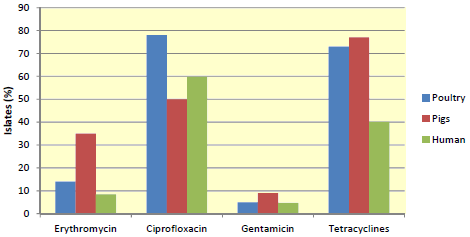
The resistance patterns in humans, especially to the macrolide, erythromycin, are more similar to poultry than pig. This can be explained by the fact that most broiler carcasses are infected with both C. jejuni and C. coli and they are primarily responsible for the infection in Man. Treatment of pig carcasses (cooling etc) helps reduce the contamination level almost to zero and the risk of transmission of C. coli is therefore very low (Burch, 2002). Of interest, fluoroquinolone resistance is higher than for S. Typhimurium as it is a one-step mutation that leads to full resistance in Campylobacter spp.
Conclusions
There are some major issues concerning antimicrobial use and resistance development and the possible spread of these resistant organisms or genes to Man. MRSA has been a real wake-up warning to a number of factors. How we use cephalosporins in pigs and other species but also the trading routes for pigs and poultry and the spread of resistant clones across Europe. Monitoring of ESBL resistance in E. coli in Europe has identified that the risk from pigs is relatively small but in chickens it is much higher. This is of concern as cephalosporins are not approved for use in chickens in the EU. This should be the first point of control. It also demonstrates that high resistance to other antibiotics has little impact on ESBL resistance development, a point ignored by the Dutch control proposals.
With regard to zoonotic infections and resistance transfer to Man, pigs do appear to play a role in S. Typhimurium transmission but from an ESBL perspective, the risk is very small. Regarding Campylobacter coli, the risk of fluoroquinolone resistance transmission would appear to be high but in fact, it is very low as few organisms are transmitted via pig meat. Poultry meat, which is the major transmitter of C. jejuni, is also the major risk for C. coli. Possibly, it is a more urgent area for control and improvement.
Responsible use of antimicrobials must be the way forward for veterinary medicine. Reduce overall use by all means, it will improve the clinical antimicrobial resistance situation and improve therapy. Improve management and housing and use vaccines more. Movement of animals and biosecurity on a farm and international basis would also appear to be of major significance. Restricting use of antimicrobial products in accordance with their SPCs (summary of product characteristics) is important, i.e. do not use cephalosporins as frontline drugs or use them off label but reserve them for when other drugs have failed.
There are many ways in which we can improve animal health and reduce antimicrobial use and target them in a better way. However, one thing is for sure, if we do not do put our house in order, the legislators/regulators will and not necessarily in a scientific way, that will help pig production.
This article was first published in 'Suis'.
February 2012








